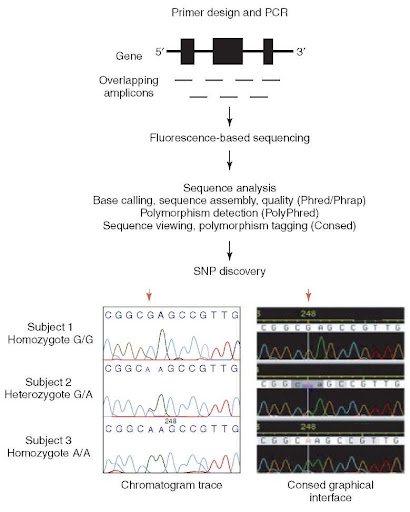
Conventional cloning of Fab and scFv libraries use multiple PCRs to introduce diversity into CDR regions and then an overlapping extension PCR to amplify the scFv or separate VL and VH chains for Fabs. Synthetic antibody libraries are usually based on a single or very low number of “scaffold” sequences with diversity introduced into one or more of the complementarity determining regions (CDRs). VHHs, also known as single-domain antibodies or nanobodies, are derived from natural heavy chain antibodies found in camelids that lack a light chain. ScFvs consist of just the VH and VL chains joined together by a peptide linker with one chain fused to the pIII protein, and VHHs consist of a single heavy chain domain equivalent to a conventional IgG VH domain. In terms of antibody formats that are amenable to phage display, Fabs consist of the VH–CH1 and VL–CL chains, one of which is fused to the pIII protein. have described a methodology for the production of high-diversity peptide libraries using a whole-plasmid PCR self-ligation process which vastly simplifies and increases the efficiency of library construction. The circularised vector, containing a short single-stranded gap, is then used for bacterial transformation. Alternatively, short oligonucleotides can be annealed to either side of the region of diversity encoded within a longer oligonucleotide such that they form sticky ends compatible with direct ligation into restriction enzyme cut vector. Synthetic peptide libraries are usually cloned by the annealing of overlapping complementary oligonucleotides, one of which codes for the peptide diversity, followed by complementary strand synthesis and digestion of the products with restriction enzymes to facilitate cloning into the phage or phagemid vector. It may therefore be required to produce bespoke synthetic one-pot ligand libraries containing a high diversity of potential binders that can provide antibodies or peptides to any target antigen with no constraints on use. Whilst there are commercial sources of both peptide and antibody libraries, the exploitation of any isolated ligand is often constrained by ownership of the libraries. 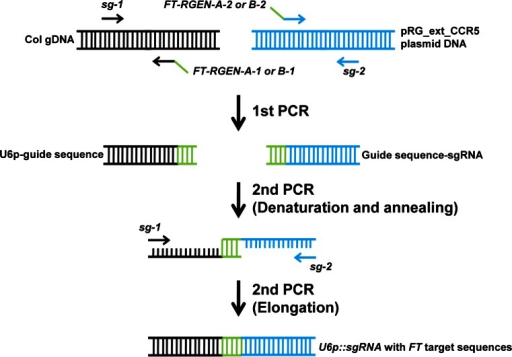
Phage display libraries can enable the identification of a variety of ligands with desirable binding properties including ligands that bind to toxic or non-immunogenic antigens. In general, relatively small ligands such as peptides are usually displayed on either coat protein pIII or pVIII and larger protein fragments such as those derived from antibodies are usually displayed on pIII. These phage libraries can display vast diversities of ligands on coat proteins projecting from the surface of the bacteriophage particle, utilising either a phagemid or phage vector system to encode the ligand-coat protein fusion. Phage display technology was first described in 1985 by George Smith and has been developed to include the display of peptides, various recombinant antibody formats, enzymes and fragmented proteomes. The VHH libraries were pooled and used to select an antibody to recombinant human collagen type 1. The peptide library was used to epitope map a monoclonal antibody. The functionality of both peptide and antibody libraries were demonstrated by selection of ligands with specific binding properties by biopanning. This method reproducibly produced 1 × 10 9 variants from just 10 transformations and the four libraries had relatively low bias with 82 to 86% of all sequences present as single copies. The cloning strategy used a simple whole-plasmid PCR method and type IIS restriction enzyme assembly that facilitate the seamless insertion of diversity into any suitable phage coat protein or antibody scaffold. Here, a peptide library of ~ 1 × 10 9 variants for display on gene VIII was produced alongside three VHH antibody libraries with similar diversity, where 12mer, 16mer or 21mer CDR3s were introduced into the highly stable cAbBCII10 scaffold displayed on gene III. The production of such libraries can be labour intensive and technically challenging and whilst there are commercial sources of libraries, the exploitation of the resulting binders is constrained by ownership of the libraries. Phage display technology utilises peptide and antibody libraries with very high diversities to select ligands with specific binding properties.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
